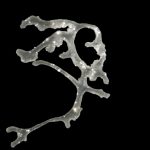
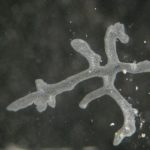
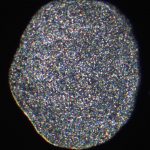
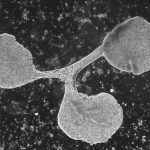
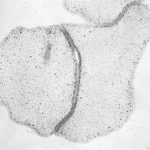
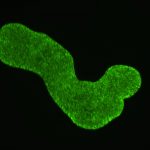
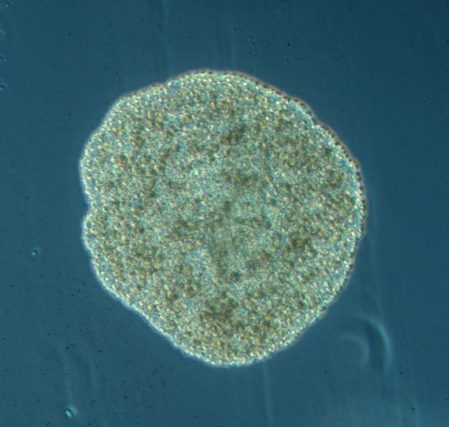
The most simple metazoan animals, the placozoans, have attracted substantial attention in different areas of biological and lately also biomedical research. The placozoan nuclear genome is the smallest – not secondarily reduced – genome among metazoan animals, yet it harbors all principal gene families known from higher animals. The phylum Placozoa has been monotypic with the single species, Trichoplax adhaerens (Schulze 1883), for more than a century. Only very recently we described two new species, with another more than 20 species collected in temperate to tropical marine waters across the globe awaiting their formal description. The yet missing – and cum grano salis perfect tabula rasa – systematics of the phylum Placozoa is unique in the animal Tree of Life.
Over the last 30 years our original Trichoplax Grell clone has been cultured meanwhile for almost 90 years in Germany and it is still going strong. We have provided Trichoplax to other labs about 130 times over the last 30 years.
We have been developing the most simple – and possibly oldest – metazoan animals as a model system for basic research across disciplines. It has turned out that placozoans represent promising – and often ideal – model systems for developing new species and systematics concepts, addressing phylogenomics, studying epithelia cell organization, unraveling ancestral features of animal physiology and development and also to address the genetics behind apoptosis and cancer. By studying organismal functions in an evolutionary context and by sequencing the genome and transcriptome of all species in the phylum we seek to contribute to a broader use of placozoans in basic and applied scientific research.
Please stop by at our lab and take a look at these fascinating animals!
DigitalMarine educational and research training
As part of the EU-funded, state-of-the-art DigitalMarine research and teaching activities we have been collecting the closest living relative of the “urmetazoon”, the placozoan Trichoplax adhaerens, from different oceans around the world. Right at the foot of the birthplace of marine biology in Europe, Bernd Schierwater has been collecting placozoans barehanded and with his jeans on since 1985.
Among many highlights for the students is the new ABC (Aerospace, Biology, Cancer) model system, the placozoan Trichoplax adhaerens. These most primitive of all metazoan animals can be collected from warm ocean waters with your pants on (see photo). Among many model system applications the ABC use of Trichoplax in a synergy of space research and developmental genetics in order to fight cancer in humans is one that fascinates not only students.
Current Collaborators
Gisella Caccone (Yale New Haven, US), Rob DeSalle (AMNH, New York, USA), Michael Eitel (LMU München, Germany), Jens Hauslage & Ruth Hemmersbach (DLR Cologne, Germany), Patrick Humbert (La Trobe Melbourne, Australia), Marc Kvansakul (La Trobe Melbourne), Frederic Nowak & Iain D. Couzin (Univ. Konstanz, Germany), Andrea Pasini & Andre Le Bivic, (Univ. Marseille, France), Frédérique Varoqueux & Dirk Fasshauer (Lausanne Univ, Switzerland).
Recent Publications
- Schleicherova D, Dulias K, Osigus HJ, Paknia O, Hadrys H, Schierwater B (2017) The most primitive metazoan animals, the placozoans, show high sensitivity to increasing ocean temperatures and acidities. Ecology and Evolution 1-10, DOI: 10.1002/ece3.2678
- Rach J, Bergmann T, Paknia O, DeSalle R, Schierwater B, Hadrys H (2017) The marker choice: Unexpected resolving power of an unexplored CO1 region for layered DNA barcoding approaches. PLOS ONE 12, DOI: 10.1371/journal.pone.0174842
- DeSalle R, Schierwater B, Hadrys H (2017) MtDNA: The small workhorse of evolutionary studies. Frontiers in Bioscience Landmark 22, 873-887.
- Novotný JP, Chughtai AA, Kostrouchová M, Kostrouchová V, Kostrouch D, Kaššák F, Kaňa R, Schierwater B, Kostrouchová M, Kostrouch Z. 2017. Trichoplax adhaerens reveals a network of nuclear receptors sensitive to 9-cis-retinoic acid at the base of metazoan evolution. PeerJ 5:e3789, DOI: 10.7717/peerj.3789
- Osigus HJ, Eitel M, Schierwater B (2017) Deep RNA sequencing reveals the smallest known and smallest possible micro exon: The placozoan cox1 single base pair exon, PLOS ONE, DOI: 10.1371/journal.pone.0177959
- Eitel M, Francis W, Varoqueaux F, Daraspe J, Osigus HJ, Krebs S, Vargas S, Blum H, Williams G, Schierwater B, Wörheide G (2018) Compartative genomics and the nature of placozoan species PLoS Biology 16(7), DOI: 10.137/journal.pbio.2005359.
- Gul IS, Staal J, Hulpiau P, de Keuckelaere E, Kamm K, Deroo T, Sanders E, Staes K, Driege Y, Saeys Y, Beyaert R, Technau U, Schierwater B and van Roy F (2018) GC content of early metazoan genes and its impact on gene expression levels in mammalian cell lines. Genome Biol Evol 10(3), 909-917, DOI: 10.1093/gbe/evy040.
- Schierwater B, DeSalle R (2018) Placozoa. Curr Biol 28, R 89-102.
- Roth SK, Powell A, Smith DJ, Roth F, Schierwater B (2018). The highly competitive ascidian Didemnum sp. threatens coral reef communities in the Wakatobi Marine National Park, Southeast Sulawesi, Indonesia. Reg Stud Mar Sci 24, 48-54, DOI: 10.1016/j.rsma.2018.07.001
- Kamm K, Osigus HJ, Stadler P, DeSalle R, Schierwater B (2018) Trichoplax Genomes Reveal New Mode of Speciation in Placozoa. Sci Rep 8, 11168, DOI: 10.1038/s41598-018-29400-y
- Varoqueaux F, Williams E A, Grandemange S, Truscello L, Kamm K, Schierwater B, Jékely G, and Fasshauer D (2018). High Cell Diversity and Complex Peptidergic Signaling Underlie Placozoan Behavior. Curr Biol. 2018;28(21):3495‐3501.e2. DOI: 10.1016/j.cub.2018.08.067
- Nanda S, Mahapatra S, Lindeen S, Bernau J, Cutshall S, Schierwater B, Chon T, Wahner-Roedler D, Bauer B (2019) Evaluation of a Novel Wellness Assessment Device (Preventiometer): A feasibility Pilot Study. Glob Adv Health Med, DOI: 10.1177/2164956119881096
- Kamm K, Schierwater B, and DeSalle R (2019) Innate immunity in the simplest animals – placozoans. BMC Genomics. 2019; 20: 5. DOI: 10.1186/s12864-018-5377-3
- Arvanitidis CD et al. (2018) Research Infrastructures offer capacity to address scientific questions never attempted before: Are all taxa equal? PeerJPreprints. DOI: 10.7287/peerj.preprints.26819v2
- Osigus HJ, Rolfes S, Herzog R, Kamm K, Schierwater B (2019) Polyplacotoma mediterranea is a new ramified placozoan species. Current Biology, DOI: 10.1016/j.cub.2019.01.068
- Kamm, K, Osigus, HJ, Stadler, PF, DeSalle, R & Schierwater B (2019) Genome analyses of a placozoan rickettsial endosymbiont show a combination of mutualistic and parasitic traits. Sci. Rep. 9, 17561. DOI: 10.1038/s41598-019-54037-w
- Neumann J, DeSalle R, Narechania A, Schierwater B, Tessler M (2020) Morphological character can strongly influence early animal relationships inferred from phylogenomic datasets. Syst. Biol. DOI: 10.1093/sysbio/syaa038